How to Improve Your Search for Reagents

How to Improve Your Search for Reagents
What researchers can learn from our experience with reagent data tools
On the hunt for a new antibody or antigen?
It’s likely that your pursuit will either start with or involve a search engine throughout the process. This trend has become the preferred approach as researchers are often unaware of the disadvantages involved. Search technologies have greatly improved in recent times, but they aren't sophisticated enough to effectively present complex biomaterial such as reagents...yet.
Having developed a reagent-specific search tool, the late kbFINDER; we gained direct exposure to the back-end requirements and functional limitations of search technology. The experience provided us with the following insight that every researcher should consider…
Counter-productivity
It became increasingly evident that utilizing search tools to find a specific reagent wasn’t exactly reliable and was, in some cases, misleading to researchers. Often recognized by search result messages like “no results found” or “1,639 products match your query” ; varying search tools can lead you down different rabbit holes, when you are simply trying to find a target protein with a specific feature or data. Subsequently, when you do find a potential candidate, the reagent data is in a datasheet you need to request. Next thing you know, you’ve downloaded multiple pdf files that you need to sort and compare… which may not even match your needs. The underlying issues that lead to these situations are not intentional, but rather a consequence of some of the limitations we have dissected below.
Data Consistency
The challenges of getting bio-reagents to play nice with search technology go beyond a simple IT (Information Technology) problem. These challenges largely stem from the high complexity of bio-reagent data. Getting this data to fit within a structured database can very difficult, and many organizations find themselves making compromises either by simplifying the data, or dealing with an unwieldy database that becomes ever more difficult to work with as the needs of the organization change, or as they add new reagents and target proteins in batches with customized features.
With a lot of forethought, planning, and properly leveraging data relationships - bio-reagent data can be standardized within a functional and flexible database, but this is rare, and never translates across different organizations. The case of organizations regularly failing to properly structure their bio-reagent data within a database has resulted in consistently unfriendly user experiences, and the development of user fatigue among scientists.
E-commerce Centricity
Online shopping or “e-commerce” is ultimately designed to get you from the search bar to the shopping cart. Very little focus has been invested in the qualifying process in-between. This is because most products dominating this space are commodities. As much as we would love for antibodies, antigens and enzymes to be commodities, they simply are not. They are highly specific, and the closer they relate to your experiments' design, the more you'll come to learn this (the hard way).
The reality is that before e-commerce, all these reagents were sold via distributors and vendors who piled a bunch of manufacturers products onto their cluttered catalogs, resulting in a sales rep saving the day with their process. Today, the internet empowers the manufacturer, and we are seeing them become more exclusive and in control of their products. One might argue that manufacturers trusting their products with distributors enabled the commodification of reagents, but that is not relevant to the topic at hand. The point is that all these print catalogs were converted to online catalogs, and their manually copy/pasted data populate standard searches (Google, Bing. etc.). This means search results regularly list two or more identical products from different vendors (including the manufacturers) and they would all lead to a page that was built around making the end-user hit “add to cart”. This is now dramatically changing (especially on Google). The search giants recognize these issues of data overlap and redundancy, and are increasingly omitting e-commerce pages from their results. Results are now based on scoring valuable content. Which is great, except now the researcher is at the mercy of the companies with the best search engine optimization (SEO) strategy. The best content for reagents are their citations, application data and technical literature. Which happen to be indistinguishable from other forms of content. Other (repeat) content can easily populate the first few pages of results, when the right protein can actually be found in the deeper pages.
In the end-relational databases, SEO and other approaches are essentially workarounds with an unchanged goal; prioritize a product or page in search to initiate an “add to cart”. In other words, they are solutions to problems the seller faces. They are not focused on solving for any problems of the end-user or researcher.
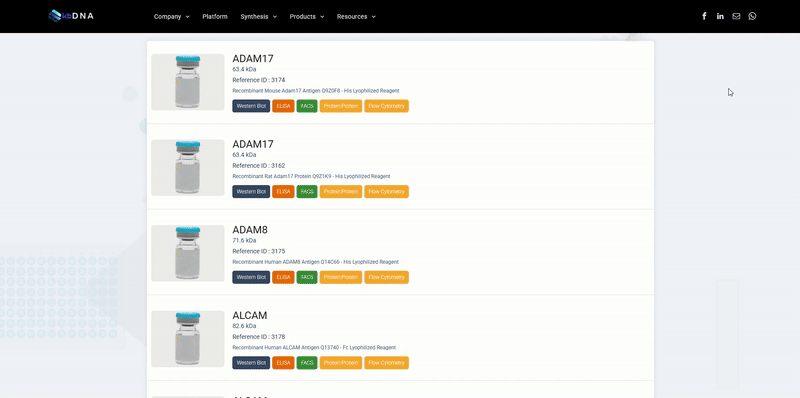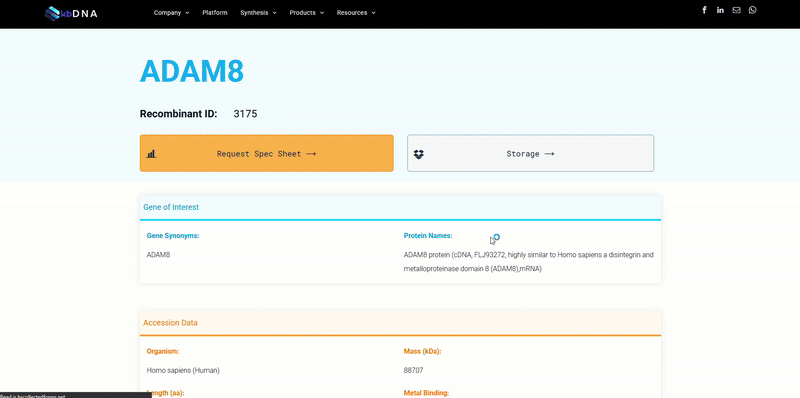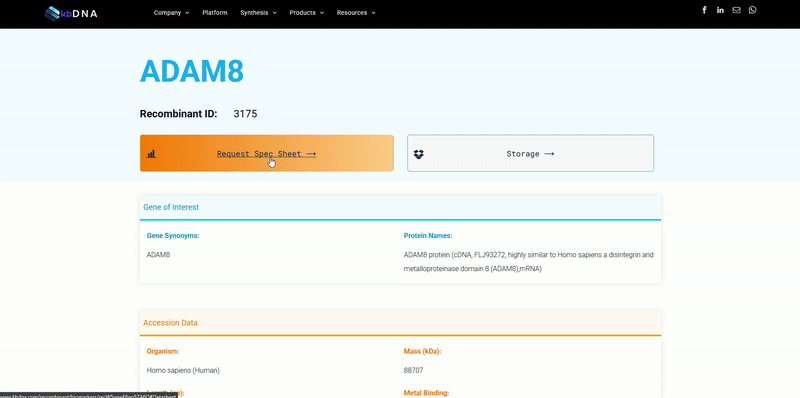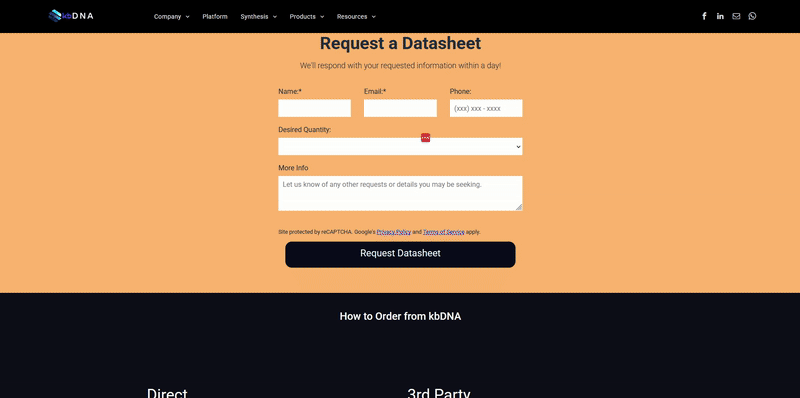
Not too long ago, our website search bar looked like this….
Now, search and e-commerce features are completely gone from the platform. After multiple releases of improved versions, kbFINDER was a very precise search tool, reinforced with a consistent and elaborate database. The more we advanced it, the more we realized that it was just another workaround the real problems.
We’ve measured greater success matching researchers with the right protein since the introduction of our recombinant libraries. The short-term benefits and long-term potential for AI compatibility make it an exciting novel solution to consider. However, libraries as an alternative to search is a challenge that will take time and effort to build correctly. Regardless of what the end-user prefers, we encourage researchers to develop a sourcing strategy that involves multiple tools/resources and to always measure the data against the standard databases (uniprot, refseq, etc.). The primary objective now is to use this insight to your advantage. Consider the outlined points in your next reagent hunt and see if you can identify any new errors!