The Synthetic Dogma

The Synthetic Dogma
How nature's molecules flow in bio-research
Introduction
Many are familiar with the central dogma of molecular biology, which demonstrates the flow of genetic material; DNA-> RNA-> Protein. However, the movement of these biomolecules as research tools requires a bit more elaboration. Fascinating as they are complex; using biomolecules to study the data, encompassing genetic material has forced the development of synthetic imitation tools. These tools are often referred to as “reagents” (or “bioreagents”). To help represent the flow of these synthetic biomolecules, kbDNA introduces its synthetic dogma.
Our synthetic dogma demonstrates the profiles of DNA, RNA & Protein as synthetic reagents and their roles throughout the flow of applications that help research genetic material:
cDNA | ssDNA
DNA is the traditional focus when it comes to genetic material.
Molecular cloning helped put genes in the hands of biologists with the production of complementary DNA (cDNA) clones. Popular formats for cDNA reagents are full length, open reading frame (ORF), 3’ untranslated region (UTR) and mutant clones. Each option designed to support specific applications.
Synthetic DNA can also be achieved by way of assembly. Using basepair fragments, called gBlocks; double-stranded A,C,T,G fragments can be chained together into a target DNA sequence. The method of choice for this technology is the infamous Gibson Assembly Protocol. gBlock DNA has contributed significant improvements in the application of DNA libraries for both molecular analysis and commercial product development.
Single-stranded DNA (ssDNA) also plays a key role in genetic studies such as expression analysis. ssDNA is part of the oligonucleotide (oligo) family and derives from the methods of solid-phase synthesis. They help target the short-sequence capacity of DNA rather than functional genes. ssDNA sequences are often customized to fit a wide range of applications. Their use can be as simple as being primers for PCR or as difficult as libraries for biosensors.
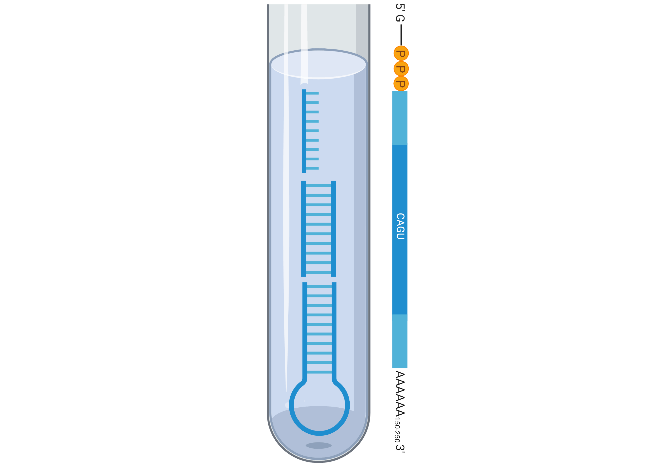


ssRNA | dsRNA
RNA is quickly climbing as the solution to research design.
Available in its various forms, RNA has been a key player in understanding the bridge between genetics and proteomics. Traditional focus has centered around its translational properties as messenger RNA (mRNA) and a good deal of mRNA reagents were developed to support an added layer of gene expression analysis. mRNA is part of the single-stranded RNA (ssRNA) oligo family, making it a benchmark in solid-phase synthesis and synthetic biology.
Most recently, RNA has taken a larger position in both experimental analysis and therapeutic design. In contrast to ssRNA, double-stranded RNA (dsRNA) synthesis showcases the advancing structural capabilities in therapeutic design. More specifically in antisense technology or “RNAi”. RNAi involves a collection of both ssRNA and dsRNA variations including; micro (miRNA), small interfering (siRNA), short hairpin (shRNA) designed to carry out the RNAi silencing mechanisms. With improving chemical compounds, RNA is matched with extensive modifications, continuing to extend its influence into more technology and applications. Two great examples are guide RNA (gRNA) in gene-editing and lipid nanoparticles (LNP-RNA) in transfection/drug delivery.
Recombinant Protein | Peptides
Protein is at the center of all bioscientific research.
Most genes are protein-coding DNA. Meaning their objective is to express a specific peptide or protein of interest-specific to its function. Protein reagents can be simplified into 3 categories; antibody, antigen, functional/enzyme. Each which have their own extensive list of subcategories based on their structure, pathway, etc. In short; antibodies detect, antigens are detected, functional/enzymes perform a target mechanism. Although it is definitely not that simple, they are all produced by way of recombination. Recombination systems use a variety of organisms (bacterial, mammalian, yeast, insect) cells to chemically express DNA (often cDNA) into a protein. This now recombinant protein is then used in a long list of applications; immunoblotting, flow cytometry, crystallography and of course assays, assays, assays. All which collect very complex data to support and/or validate the research.
All roads lead to protein and that responsibility comes with its own list of complications. Not only are protein structures very difficult to achieve beyond a certain threshold, but their precise functional data is very laborious to collect. These issues have created bottleneck-like limitations in assay development and relational proteomics. While methods such as cell-free expression systems build production solutions, the road to bioinformatics shows promise to organizing both commercial and research protein data.
kbDNA, INC.
125 Cambridgepark Dr.
Cambridge, MA 02140
Company
Contact Us
Phone:
+1 (781) 206-2235
Fax:
+1 (781) 206-2258
Email:
info@kbDNA.com
Useful Links
The kbDNA Inc. Quality Management Network is certified as conforming to ISO 9001:2015 standard. Request Certificate